About iRNA-m5C
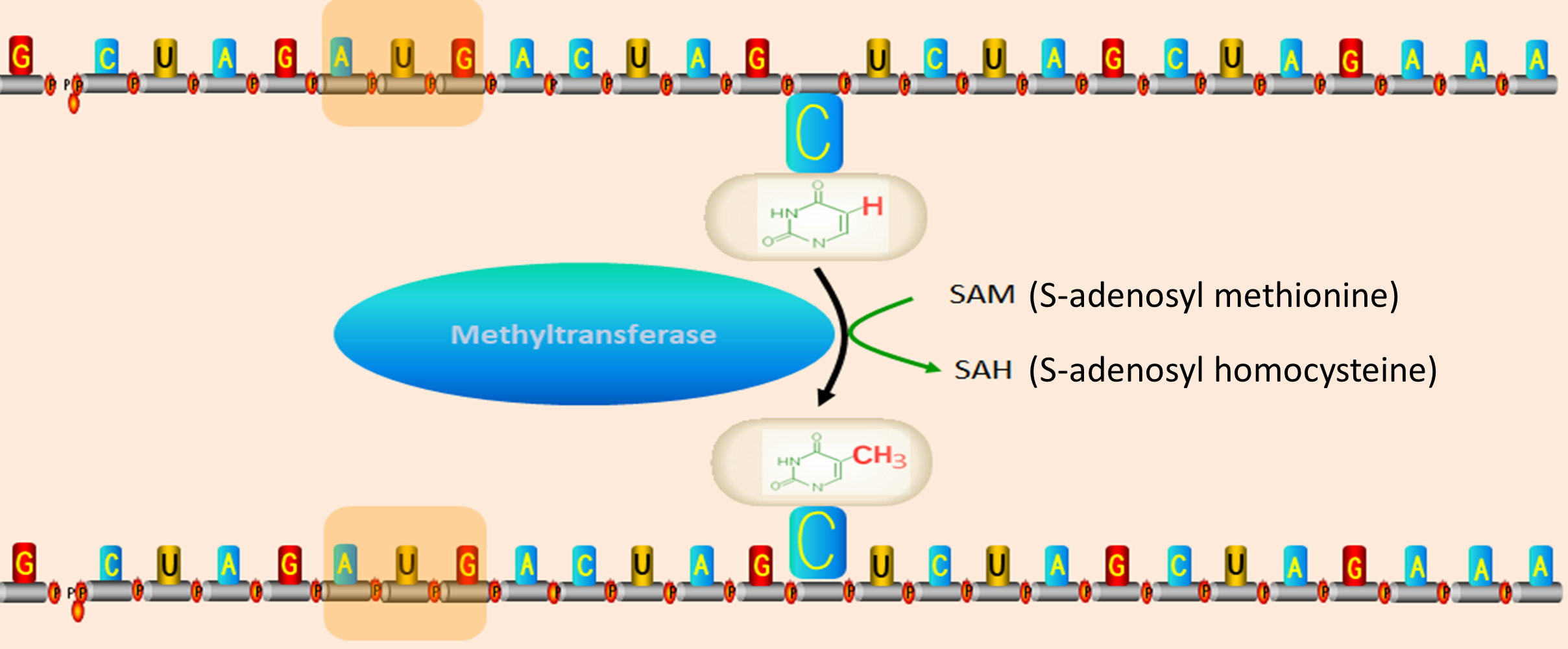
5-methylcytosine (m5C) is formed by transferring methyl groups to the 5-th position of the cytosine ring catalyzed by RNA methyltransferases. Accurate identification of m5C site is a key step in understanding its biological functions. In this work, we constructed a more powerful and reliable model for identifying m5C sites. To train the model, we collected experimentally confirmed m5C data from Homo Sapiens (H. Sapiens), Mus musculus (M. musculus), Saccharomyces cerevisiae (S. cerevisiae) and Arabidopsis thaliana (A. thaliana), and compared the performances of different feature extraction methods and classification algorithms for optimizing prediction model. Based on the optimal model, a novel predictor called iRNA-m5C was developed for the recognition of m5C sites. Finally, we critically evaluated the performance of iRNA-m5C and compared it with existing methods. The result showed that iRNA-m5C could produce the best prediction performance. We anticipate that the iRNA-m5C will become a powerful tool for large scale identification of m5C sites.